The Non-dominant AAA+ Ring in the ClpAP Protease Functions as an Anti-stalling Motor to Accelerate Protein Unfolding and Translocation - ScienceDirect
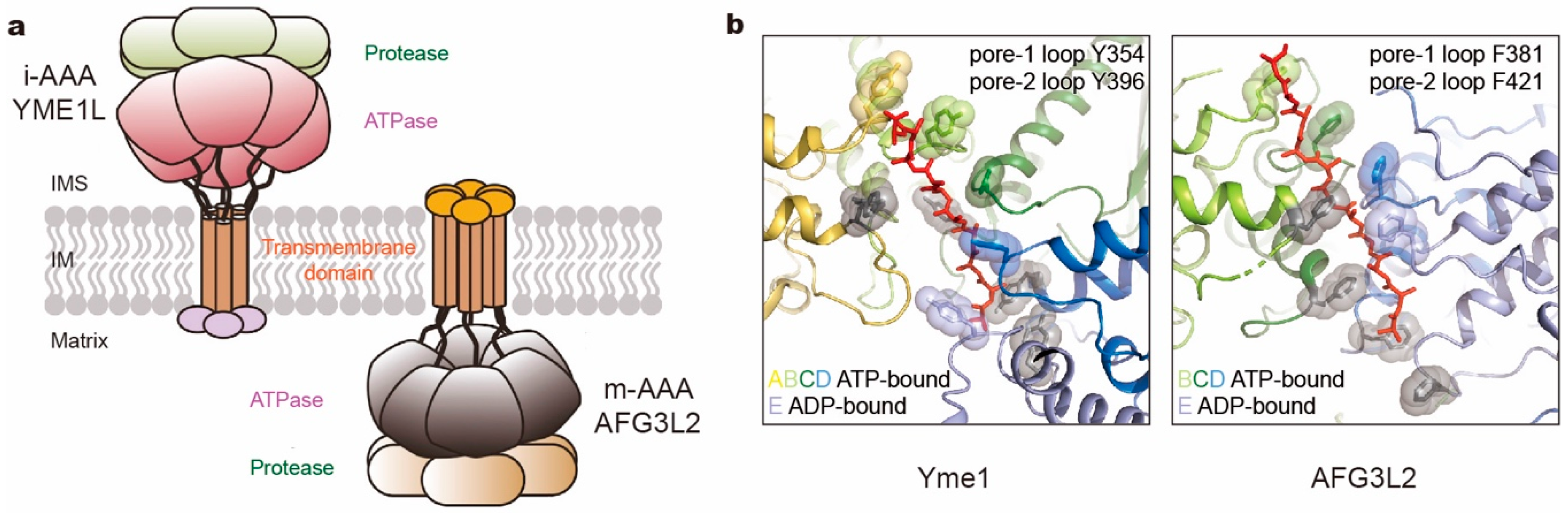
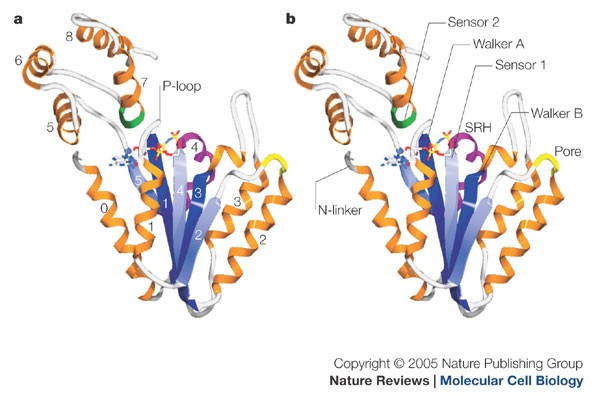
Biomolecules | Free Full-Text | AAA+ ATPases in Protein Degradation: Structures, Functions and Mechanisms
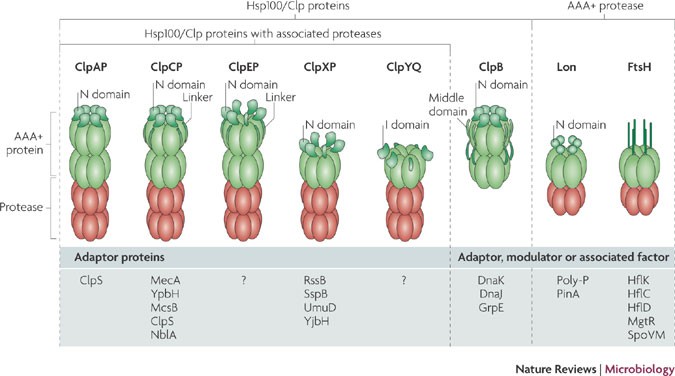
Adapting the machine: adaptor proteins for Hsp100/Clp and AAA+ proteases | Nature Reviews Microbiology

Dynamic functional assembly of the Torsin AAA+ ATPase and its modulation by LAP1 | Molecular Biology of the Cell

Biomolecules | Free Full-Text | AAA+ ATPases in Protein Degradation: Structures, Functions and Mechanisms

Asymmetric Interactions of ATP with the AAA+ ClpX6 Unfoldase: Allosteric Control of a Protein Machine: Cell

MuB is an AAA+ ATPase that forms helical filaments to control target selection for DNA transposition | PNAS









![PDF] Structure and mechanism of the ESCRT pathway AAA+ ATPase Vps4 | Semantic Scholar PDF] Structure and mechanism of the ESCRT pathway AAA+ ATPase Vps4 | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/bdce9422f5d35c390af869af721175a3d96d5efd/2-Figure1-1.png)